Introduction
Ranaviruses from the viral family
Iridoviridae are large double-stranded DNA viruses with a large vertebrate host range (
Duffus et al. 2015;
Chinchar et al. 2017). They were first described in North American northern leopard frogs (
Rana pipiens now
Lithobates pipiens) in the mid-1960s (
Granoff et al. 1965). Since then, they have been described in over a hundred other species of amphibians, reptiles, and fish from all around the globe (
Duffus et al. 2015). Ranaviruses are responsible for countless morbidity and mortality events in affected species and have even caused population declines (e.g., common frogs (
Rana temporaria) in the UK;
Teacher et al. 2010) and local extirpations (e.g., multiple species in the Spanish Pyrenees;
Price et al. 2014). They are also known to affect threatened and endangered species such as the Chinese giant salamander (
Andrias davidanus,
Geng et al. 2011) and pallid sturgeon (
Scaphirhynchus albus,
Waltzek et al. 2014), making them a cause for conservation concern.
Currently, there are six species of ranavirus recognized by the International Committee on Taxonomy of Viruses (ICTV;
Chinchar et al. 2017); this is as a result of a deeper understanding of
Ranavirus genomics. Initially, it was thought that the major capsid protein (MCP) was adequate for the study of viral evolution of iridoviruses, as it has both highly conserved, but also some variable domains (
Tidona et al. 1998). However, some studies have used only portions of the MCP and have had trouble parsing possible local adaptations (e.g.,
Duffus and Andrews 2013). Other studies have used multiple genes, including the MCP, and have been able to detect genetic variation in large-scale data sets (e.g.,
Stöhr et al. 2015) and in genetic information from geographically isolated data sets (
Ridenhour and Storfer 2008). While the MCP gene may have enough variation to place ranaviruses into their different species, a study that uses only the MCP may miss local variation or adaptation. For example,
Ridenhour and Storfer (2008) only found evidence of local adaptation of
Ambystoma tigrinum virus strains in the southeastern United States when they examined >5% of the genome and included genes such as the eIF-2α homologue, which is not present in all iridoviruses.
Eaton et al. (2007) reexamined the genomes of 12 different iridoviruses. They found that there were 26 core genes across all iridoviruses. These core genes are typically conserved, with functions that are generally associated with virulence, replication, and gene expression (
Eaton et al. 2007;
Jancovich et al. 2015). However, further molecular studies of newly discovered and sequenced iridovirids bring this total of shared genes down to 24 (e.g., Shrimp hemocyte iridescent virus lacks the small subunit of ribonucleotide reductase (
Qiu et al. 2018) and European Chub Iridovirus lacks a deoxinucleotide reductase (GenBank accession number MK6376310)). Here we examine the utility of 24 core iridovirus genes across six species of ranavirus and perform a large scale dot-plot analysis of 17 different isolates. However, it is important to note that all ranaviruses contain the original 26 core genes and a number of other genes that are found in all ranaviruses. We hope that the results from this study will be useful for those who study ranavirus phylogenomics, genomics, and phylogenetics.
Results
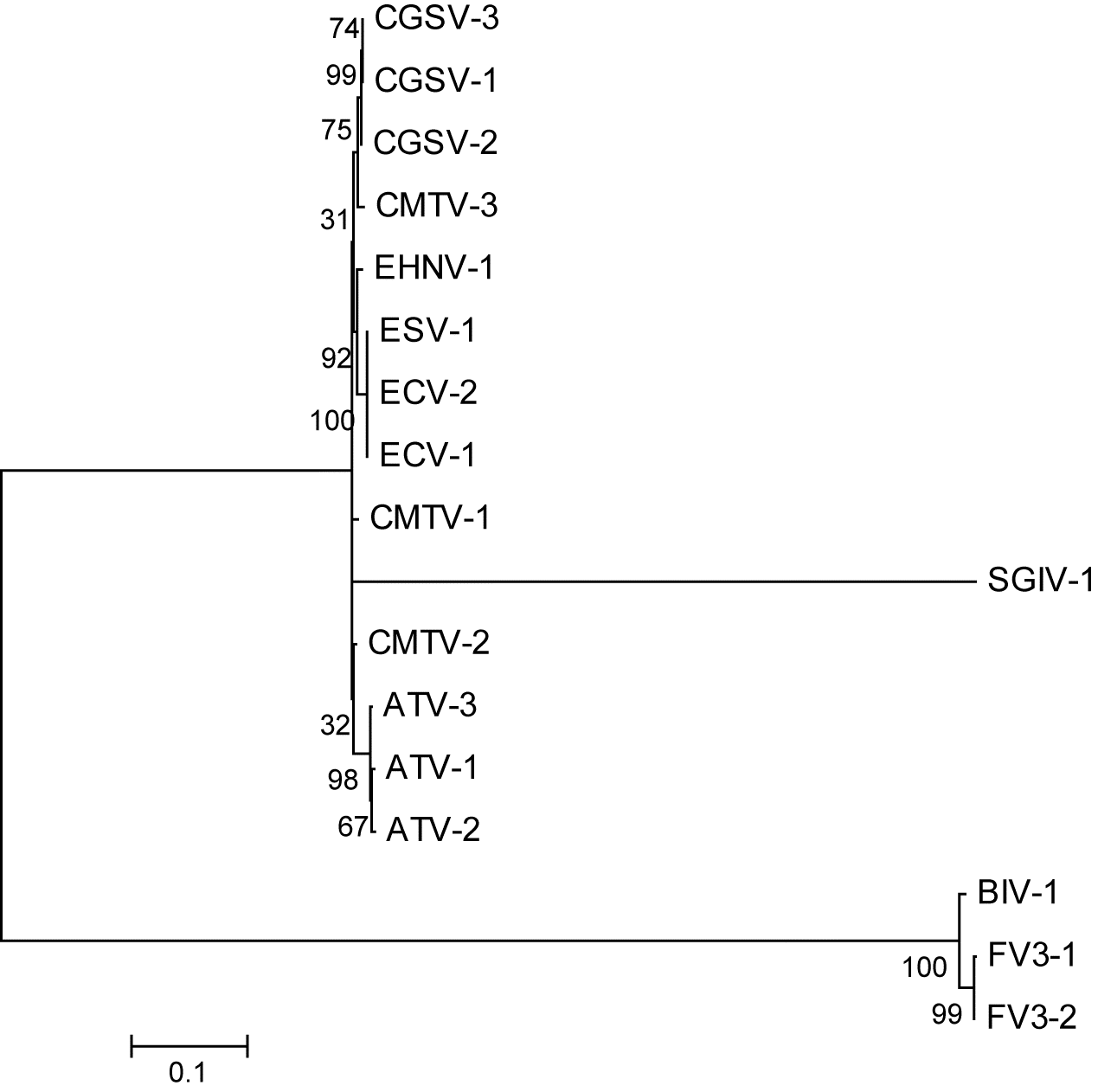
Most of the phylogenetic trees for the 24 core genes of the 17 isolates of ranavirus that were examined were able to group at the species level (six species). SGIV, however, is problematic as it appears as an individual group. It is important to note that at the time of this analysis only one sequence of SGIV was available; however, Grouper Iridovirus (not included in this analysis) may be a strain of SGIV. The tree constructed with full genome sequences (
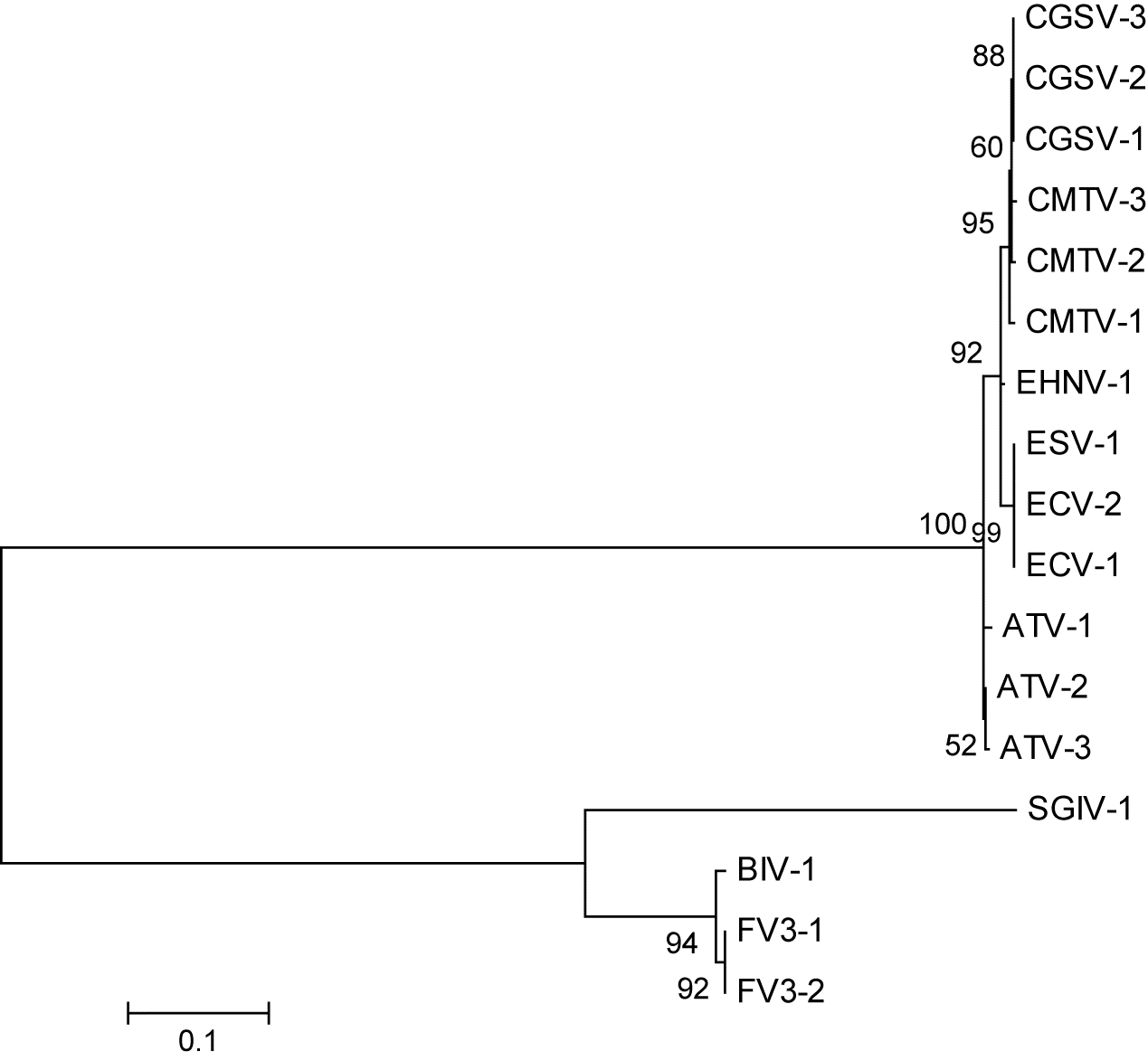
Fig. 1) shows the usual groupings of the six ranavirus species used in this study. Visually, the best trees, besides those made by the major capsid protein (ORF 90R in
Frog virus 3 (FV3);
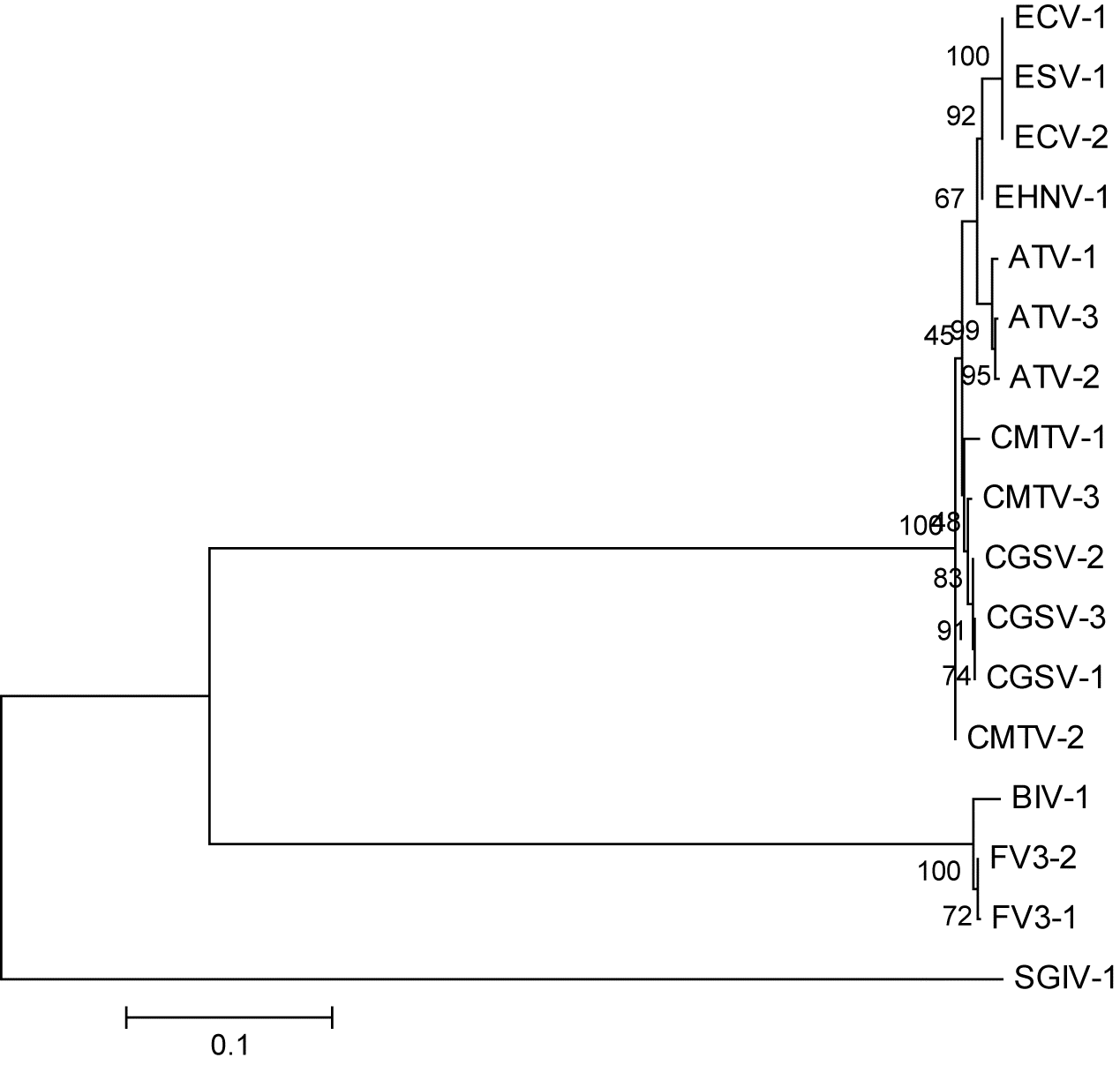
Fig. 2), are ORF 19R in FV3 (
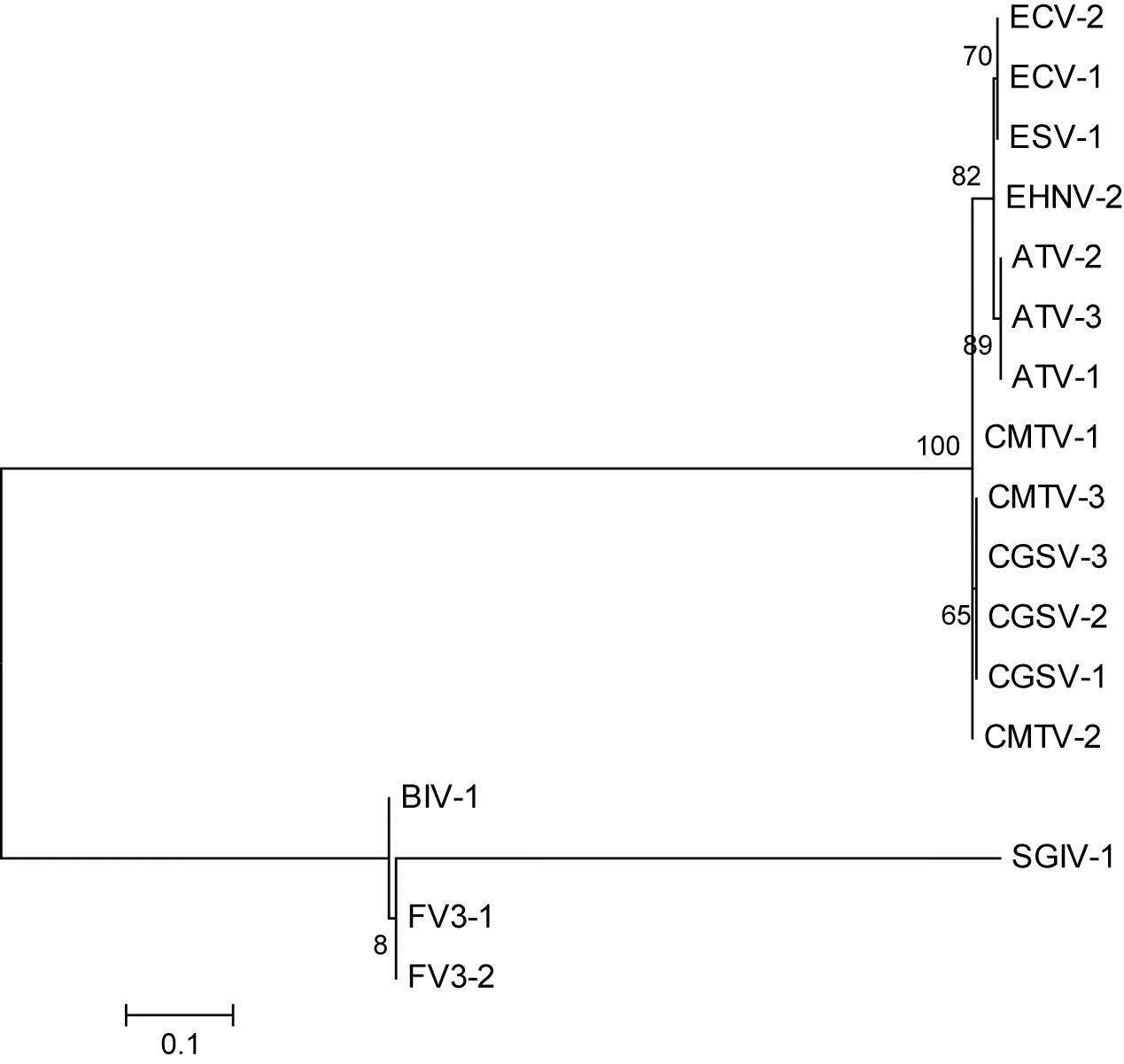
Fig. 3), and ORF 88R in FV3 (
Fig. 4). One of the worst ORFs for reconstruction was ORF 1R in FV3 (
Fig. S1) because the branch lengths were extremely short between the different species with low bootstrap support. (All other trees can be found in the
Figs. S2–
S22.)
The trees that had the shortest symmetrical distance (i.e., 10 or below) to the MCP were made from ORFs 12L, 19R, 60R, 62L, 67L, 80L, 81R, 88R, 91R, and 94L. However, trees that had the shortest symmetrical distance to the full genomes were ORF 91R, ORF 94L, and ORF 95R (Table 2).
The best trees over all were made from ORF 19R, ORF 88R, ORF 91R (
Fig. S19), and ORF 94L (
Fig. S20) when all three types of analyses were considered together.
Dot-plot analysis of all 17 isolates of ranavirus used in the current study showed collinearity at the species level (e.g., Common midwife toad virus and Chinese Giant Salamander Virus, FV3, and Bohle Iridovirus) and there tends to only be smaller genomic rearrangements between most of the different species of
Ranavirus. However, SGIV is not collinear with any other ranaviruses analyzed in this study (see
Fig. 5).
Discussion
While most of the 24 core iridovirus genes used in this study will sort out the different ranavirus isolates to the species level, their utility is quite limited for finer-scale differentiation. There are only two genes that we feel make good approximations to phylogenetic trees constructed with the whole genome or the full sequence of the major capsid protein gene (ORF 90R in FV3), these are ORFs 19R and 88R when visually compared. ORF 19R is hypothesized to be a serine–threonine protein kinase (
Eaton et al. 2007). This group of enzymes typically catalyze the transfer of phosphate groups from adenosine triphosphate (ATP) to proteins (
Jacob et al. 2011). Our results here are surprising because viral serine–threonine protein kinases are usually highly conserved (
Jacob et al. 2011). However, iridoviruses are hypothesized to have two serine–threonine protein kinases (ORF57R is a second FV3 serine–threonine kinase) in their core gene set (
Eaton et al. 2007). It is common for genes that are duplicated in the genome to have one that is more variable than the other because of differences in selective pressures and mutation (see
Shackelton and Holmes (2004) for a review of large DNA virus evolution). This is likely what is occurring in the case of ORFs 19R and 57R.
Open reading frame 88R is hypothesized to be an Erv I/Alr family protein (
Eaton et al. 2007). These are sulfhydryl oxidase and are they are known to be far more divergent in viruses than in other organisms (
Fass 2008). This sequence flexibility is likely what permits for a finer-scale differentiation between ranavirus species than other genes that are highly conserved based on functional needs. There is only a single gene that codes for Erv I/Alr family proteins in the core genes (
Eaton et al. 2007), leaving out the potential for a gene duplication event and subsequent divergent evolution of the two genes within the iridovirus core genome.
Open reading frame 91R is hypothesized to be an immediate early protein infected-cell protein —46 homologue (
Eaton et al. 2007). The function of ORF 91R has been partially characterized by
Penny and Brunetti (2019). During infection, its protein product can be found in the nucleus and it is hypothesized that it is involved with viral DNA transcription as well as involved in one or more early viral replication processes (
Penny and Brunetti 2019). Open reading frame 94L is hypothesized to be a homologue of a protein found in
Clostridium tetani (
Eaton et al. 2007). Its protein product localizes in the endoplasmic reticulum and are hypothesized to have a role in the viral modulation of cellular secretory pathways based on its structure (
Penny and Brunetti 2019).
Our dot-plot analysis shows similar genomic rearrangements within ranaviral species. However, markedly different genomic arrangements can be seen between different species of ranavirus. Interestingly, SGIV shows minimal collinearity with the other ranavirus species examined (
Fig. 5). The genome is very different in its arrangement, despite having the same core, and perhaps should be considered a different iridoviral genera instead of a species within the genus
Ranavirus. In our phylogenetic analyses, SGIV tends also to be divergent, which supports this claim and previous reports of SGIV being the lowest percent similarity of all ranaviruses (see
Stöhr et al. 2015).
Most of the 24 core iridovirus genes are able to more or less sort different ranavirus strains to the species level; however, it is highly likely that they are not able to show fine-scale differentiation (e.g., geographical or local adaptations), because they are not sufficiently polymorphic between species. Only two genes, ORFs 19R and 88R in FV3, make comparatively supported trees to the major capsid protein and full genomes visually. However, when the symmetrical distances are analyzed ORF 91R and 94L may be added to potentially informative sequences for phylogenetic analysis. We do recommend that single gene sequences alone not be used as there is evidence that multiple targeted concatenated genes are more useful in reconstructing the true phylogenies because of nucleotide shifts at the third position of the codon (e.g., yeasts,
Collins et al. 2005).
Our dot-plot analysis suggests that each species of ranavirus has undergone unique genome rearrangements that are consistent at the species level. However, SGIV is highly divergent and may need to be considered a separate genera within the Iridoviridae instead of a species of ranavirus. Future directions should include determining the phylogenetic signal for each of the core genes and examining the utility of the more divergent core genes in sorting out local adaptations.